Classifications:
Chemical descriptors are used to calculate and to develop methods for chemical property calculations (QSPR - quantitative structure-property relationship) or chemical activity (QSAR - quantitative structure-activity relationship) calculations. A common classification method for descriptors can be taken from ChemoInformatics textbooks and a collection of common molecular descriptors in the Handbook of molecular descriptors [LINK].
- 0D - bond counts, mol weight, atom counts
- 1D - fragment counts, H-Bond acc/don, Crippen, PSA, SMARTS
- 2D - topological descriptors (Balaban, Randic, Wiener, BCUT, kappa, chi)
- 3D - geometrical descriptors (3D WHIM, 3D autocorrelation, 3D-Morse) + surface properties + COMFA
- 4D - 3D coordinates + conformations (JCHEM conformer, CORINA, gold set, Crystaleye)
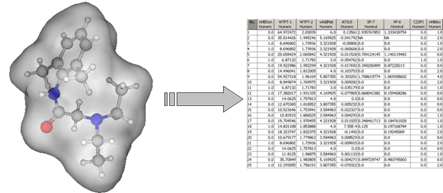
Tools for descriptor calculations:
A selection of commercial and free descriptor calculation utilities is collected under the molecular descriptor software collection or the CompChem list or new programs are posted to CCL.
- alvaDesc - new visual descriptor suite from Kode solutions covering 4000 descriptors (developed by Alvascience)
- CDK descriptor GUI (free and open source - using Open Source CDK and Joelib code)
- BlueDesc - Molecular Descriptor Calculator (free and open source - using CDK and Joelib code, requires JAVA 1.6
- ChemAxon JChem - descriptor package using Marvin JAVA API (free academic license)
- ISIDA/QSPR - free fragment based QSPR descriptor package
- E-Dragon (VCCLab) free (150 molecules), now with GSFRAG, GSFRAG-L, ETState > 3000 descriptors
- MOLD2 - (FDA) a free 2D molecule descriptor package
- Toxicity Estimation Software Tool (T.E.S.T.) - (EPA) contains more than 790 2-dimensional descriptors
- Open3DQSAR - pharmacophore modelling using molecular interaction fields (MIFs)
- Dragon - 5,270 molecular descriptors for LINUX and WIN (Todeschini/Talete/Kode)
- PaDEL-Descriptor - based on CDK but includes additional 737 2D and 3D descriptors (NUS/Singapore)
- ADMEWORKS ModelBuilder - 400 descriptors (Jurs) and MOPAC (Stewart) (Fujitsu/Poland)
- QuBiLS-MIDAS - a highly parallel software for three-dimensional molecular descriptor calculations
Concepts for descriptor calculations and QSAR/QSPR modeling:
- You need a large dataset with the molecular property (logP, bp) to be modeled. The larger the number of data points the better. There are QSAR models with 20 or less points, however for broad applications one need to cover a large diversity space. Hundreds or thousands of such values can be collected from databases or are now available from HT screening methods.
- You need the molecular structures itself (as SMILES, SDF in 2D or optimized 3D structure). Handling the molecules together with all descriptors can be a challenging task, software which can do that is highly preferred.
- You need a descriptor package for descriptor calculation
- You need to apply feature selection (a statistical process) to discard unimportant (invariant) or sometimes highly correlated descriptors (othogonalization)
- You need to divide your molecule set into three parts. A training (70%), validation (30%) and an additional external training or validation set which is not used in either method. (Sometime the validation set is called testing set or vice versa). Cross-validation (n-fold or v-fold) techniques or other resampling tests (Monte Carlo Sampling, Jackknifing, Bootstrapping) need to be applied, especially if not enough molecules are available.
- You need to apply regression or classification methods (including meta-learning approaches).
- One need to make sure that for future predictions no other compound classes are included (which usually results in wrong predictions) by either including error values, fingerprint or substructure matches or a simple dimension reduction method (PCA, PLS) to avoid molecules which were not covered during development. As example a logP method only developed on alkanes will 100% fail on complex drug molecules or molecules with multiple -OH and -NH or -SH groups. Further more a complete statistical description for either the regression performance or classification performance needs to be included.